White oak(Quercus fabri Hance)regenerated stump sprouts show few senescence symptoms during 40 years of growth in a natural forest
Honggang Sun ,Sisheng Wu and Liwen Wu
Abstract
Keywords: Sprout, Regeneration,Senescence,White oak, Vegetative propagation
Background
Sprouts are the main material of vegetative propagation,and many species, including European aspens (Populus tremula L.), Jabuticaba (Plinia jaboticaba Vell) and Populus euphratica, are propagated vegetatively through sprouts (Kramp et al. 2018; Oliveira et al. 2019; Tullus et al. 2020). Previously, it was reported that older parental trees had lower sprouting capabilities (Matula et al.2012). However, the link between parental tree age and regenerated sprout lifespan is unclear; do regenerated sprouts have shorter lifespans than sexually reproduced plants? Clonal trees are usually found to mature faster,and the growth of trees is limited by the maturation period (Greenwood 1995, Greenwood et al. 2010). This may occur because the clonal materials are mature;therefore, rejuvenation is implemented to delay or reverse maturation (Wendling et al. 2014). Ally et al.(2010) hypothesized that somatic mutations from parents are not eliminated by homologous recombination and natural selection. Consequently, ramets carry somatic mutations, which result in lower fertility rates. However, sprouts grow from calli, in which genes are reprogrammed. The telomere lengths also increase during callus culturing in species like Arabidopsis thaliana,maize, and agave (Rescalvo-Morales et al. 2016; Santos-Serejo and Aguiar-Perecin 2016; Sováková et al. 2018).Thus, the aging status may be reversed in calli. In Scots pine (Pinus sylvestris Linn.), although DNA methylation levels increase with meristematic age, the growth rates and reproductive capacities of grafted shoots do not differ between old and young donors (Mencuccini et al.2014). Owing to their long lengths, lifespans of vegetatively propagated perennial trees are hard to measure;consequently, limited studies have been published.
Accelerating senescence and leaf abscission to facilitate the survival of the next generation is a strategy of monocarpic plants (mainly annual or biennial) in response to stress, and the senescence of an entire organism occurs readily in monocarpic species (Sade et al. 2018). Unlike monocarpic plants, polycarpic plants (perennial) grow indefinitely, and plants produce new ramets to compensate for lost ones. Therefore, senescence at the organismal level rarely occurs, except when the formation of new ramets cannot offset losses (Munné-Bosch 2015).However, the accelerated senescence of leaves and roots is still harmful to polycarpic species because nutrient acquisition is curtailed. A decline in reproductive capacity is another signal of organismal senescence. White campion (Silene latifolia), a short-lived herb, produces fewer inflorescences in old age (Pujol et al. 2014), and the number of viable pollen grains per catkin per ramet decreases with clonal age in male Populus tremuloides(Ally et al. 2010). In Elymus excelsus, a bunchgrass, the reproductive capacity, but not the vegetative growth, decreases with age (Li et al., 2019a,b).
For plant organs, senescence is the final developmental stage.During this stage, several life-support systems start to decline, tissues stop growing and programmed cell death occurs on a large scale (Thomas 2013). The decline of the antioxidative system is one characteristic of senescence. Decreased antioxidase activity and antioxidant levels lead to reactive oxygen species (ROS) accumulation. ROS react with DNA and metabolites, activate several senescence-regulated NAC transcription factors,induce so-called oxidative damage and ultimately kill the cells (Berni et al. 2019; Kikuchi et al. 2020). In addition,the accumulation of gene mutations and telomere shortening are ongoing during organ life. The DNA repair system continuously upregulates to maintain genetic stability, and apoptosis occurs when DNA damage is irreparable (Abate et al. 2020). Senescence-associated genes,such as those involved in plant hormone signaling(abscisic acid-responsive element-binding transcription factors 2, 3 and 4) (Gao et al. 2016), epigenetic modifications (Arabidopsis JmjC-domain containing protein 16)(Liu et al. 2019) and pathogen defense (Senescence-associated genes 13) (Dhar et al. 2020), also play roles in senescence.
White oak (Quercus fabri Hance) is widely distributed in the northern hemisphere, and it takes more than 50 years to grow from a sapling to harvest size (Schweitzer et al. 2019). When white oaks are cut down, the stumps have a high sprouting capability (Weigel and Peng C-Y 2002). In this study, to explore whether physiological ages of parental trees influenced the lifespans of the regenerated sprouts, the senescence levels of 5-, 10-, 20-and 40-year-old white oak regenerated stump sprouts in a natural forest were evaluated and the antioxidative capabilities and transcriptomes in leaves and shoots were compared. We hypothesized that if regenerated sprouts have shorter lifespans, then the leaves and shoots of 40-year-old sprouts should show obvious signs of senescence.
Materials and methods
Study sites and plants
The study sites were located at Fengshushan Forestry Farm (29°11′N,117°32′E) in Jiangxi Province, Southern China, which experiences a subtropical monsoon climate. The mean annual(1981-2010), monthly minimum(January) and monthly maximum (July) temperatures are 17.8°C, 10.9°C and 41.8°C, respectively. The average length of the growing season is 263 days, and the mean annual precipitation is 1805 mm.All the white oaks form one population, with sprouts being regenerated from stumps and allowed to grow for 5, 10, 20 and 40 years(groups 5a, 10a, 20a and 40a, respectively). The diameters of the stumps were similar, as were the sprouts’main-stem diameters in the same age group.Fresh leaves and shoots attached to leaves on regenerated sprouts were collected in mid-July. We used pruning shears to collect leaf and shoot samples. Because the samples were collected from one population, there were only four samples in both the 5a and 40a groups (n=4) and six samples in both the 10a and 20a groups (n=6). All the leaves and shoots were stored on dry ice immediately after collection, transported to the laboratory and then stored at −80°C.
Estimates of antioxidative capabilities
A certain weight of leaf or shoot sample was crushed in a prechilled pestle and crushed with a mortar in an appropriate volume of prechilled extraction buffer. After uniform homogenization and centrifugation at 4000 g for 15 min at 4°C, the supernatant was collected and used for biochemical analyses.
Antioxidative capabilities were estimated by quantifying peroxidation products and antioxidase activity levels.The malondialdehyde (MDA) content was determined using a Plant MDA Assay Kit (TBA method; A003-3-1,Nanjing Jiancheng, Nanjing, China). The hydrogen peroxide (H2O2) level was determined using an H2O2Assay Kit (H2O2reacted with titanium salt and is then quantified at OD415; A064-1-1, Nanjing Jiancheng), and the superoxide anion (O2−) level was determined using an O2−Assay Kit (sulfanilamide method; R30342, Shanghai Yuanye, Shanghai, China). The activity levels of superoxide dismutase (SOD), peroxidase (POD), and catalase(CAT) were determined using the Total SOD (WST-1 method; A001-1-2, Nanjing Jiancheng), POD (H2O2catalysis method; A084-3-1, Nanjing Jiancheng) and CAT(H2O2catalysis method; A007-1-1, Nanjing Jiancheng)Assay Kits, respectively, following the manufacturer’s instructions.All the leaf and shoot samples were measured three times in parallel.
RNA sequencing and analysis
Total RNA was extracted using TRIzol reagent (Invitrogen, California, USA) following the manufacturer’s instructions. Quality was checked using gel electrophoresis and an Agilent 2100 Bioanalyzer (Agilent Technologies,CA, USA). Pooled libraries were sequenced on an Illumina HiSeq platform with 2×150-bp paired-end reads.After removing the adaptor and low-quality reads, the remaining clean reads were de novo assembled using Trinity software (Haas et al. 2013). Differentially expressed genes (DEGs) were determined using DESeq(R package) with the criteria fold change >2 and p-value<0.05 (Anders and Huber 2010). All the genes were annotated in Kyoto Encyclopedia of Genes and Genomes(KEGG)databases for functional classifications. The KEGG enrichment was performed using TBtools(Chen et al.2020).
The gene expression pattern analysis was performed using Short Time-series Expression Miner software(Ernst and Bar-Joseph 2006) on the OmicShare tools platform (www.omicshare.com/tools).
Statistical analysis
Data were evaluated using a one-way ANOVA followed by Tukey’s multiple comparisons tests. Values are expressed as means ± SEMs.
Results
Oxidative stress does not increase in older regenerated sprouts
Oxidative stress-related indices were different for leaves and shoots, and they were not linearly correlated with sprout ages (Fig. 1). In leaves, SOD and POD activities showed no significant changes among the four tested ages, while the CAT activities in the 10a and 40a groups were lower than in the 5a and 20a groups. The MDA,H2O2and O2−levels were lowest in the 40a group. In shoots, SOD and CAT activity levels in the 10a group were lower than in the other groups, while the POD activity in the 20a group was greater than in the 5a and 40a groups. However, the MDA, H2O2and O2−levels in shoots were not significantly different among the four tested ages. Although the enzyme activities differed among the four age groups, the transcriptional levels of the SOD, POD and CAT families were not significantly changed (Fig. 2).

Fig.1 Levels of antioxidative enzyme activities and peroxidation products in white oak leaves and shoots.SOD,superoxide dismutase; POD,peroxidase; CAT, catalase;MDA, malondialdehyde;H2O2,hydrogen peroxide,mol·g−1; O2−, superoxide anion.Different letters above columns indicate significant differences (p<0.5)
Age-related changes at the transcriptional levels
There were greater numbers of DEGs in leaves than in shoots (Fig. 3a). In leaves, the 10a:20a (L10:L20) comparison produced the lowest number of DEGs, with 185 and 183 DEGs up- and downregulated, respectively, in the L20 group. The 10a:40a (L10:L40) and 20a:40a (L20:L40) comparisons produced the greatest numbers of DEGs, with 5601 and 4094 DEGs upregulated in the L10 and L20 groups, respectively, and only 39 and 45 DEGs downregulated in the L10 and L20 groups, respectively.In total, 3124 DEGs were shared in the L10:L40 and L20:L40 comparisons, and 1613 DEGs were unique to the L10:L40 comparison (Fig. 3b). The 5a:10a comparison (L5:L10) produced 3831 DEGs, with 2006 and 1825 up- and downregulated, respectively, in the L10 group(Fig. 3a). In total, 1085 DEGs were shared in the L5:L10,L5:L20 and L5:L40 comparisons (Fig. 3b). It was apparent that main phyllosphere transcriptional changes occurred in the 5a-10a and 20a-40a stages.
In shoots, the transcriptional variation was minimal in the 5a:40a (S5:S40) comparison, with only five DEGs downregulated in the S40 group. The maximal variation occurred in the 5a:10a (S5:S10) comparison, with 1552 and 566 DEGs up- and downregulated, respectively, in the S10 group (Fig. 3a). The degrees of transcriptional variation were similar in the 10a:20a (S10:S20) and 20a:40a (S20:S40) comparisons, with 735 and 631 DEGs, respectively, and most of these DEGs were upregulated in the latter groups (Fig.3c).
Enrichment analysis reveals some unhealthy characteristics
After removing pathways with no significance (p>0.05)and those that were only distinguished in a single comparison, a KEGG enrichment analysis of DEGs indicated that several pathways changed during sprout growth(Tables 1 and 2). Programmed cell death-related pathways, including apoptosis, autophagy, and DNA mismatch repair, were not markedly upregulated in older organs (Signorelli et al. 2019).
In leaves, cyanoamino acid metabolism, flavonoid biosynthesis, phenylpropanoid biosynthesis, brassinosteroid biosynthesis, starch and sucrose metabolism, cutin, suberine and wax biosynthesis, and fatty acid elongation pathways varied across ages, showing changes in four of six comparisons (Table 1 and S1). The majority of DEGs enriched in pathways related to growth (starch and sucrose metabolism, cutin, suberine and wax biosynthesis,fatty acid elongation and RNA transport), cell division(DNA replication and both porphyrin and chlorophyll metabolism), and death (base excision repair, phagosome, proteasome and glycerophospholipid metabolism)were upregulated at greater ages(Fig. 4a).
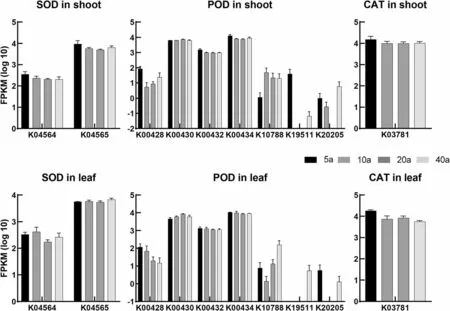
Fig.2 Transcriptional levels of the SOD,POD and CAT families in white oak leaves and shoots.FPKM,Fragments Per Kilobase of exon model per Million mapped fragments.K00518,nickel superoxide dismutase[EC:1.15.1.1];K04565,superoxide dismutase, Cu-Zn family[EC:1.15.1.1];K00428,cytochrome c peroxidase[EC:1.11.1.5];K00430,peroxidase[EC:1.11.1.7];K00432,glutathione peroxidase[EC:1.11.1.9];K00434,L-ascorbate peroxidase[EC:1.11.1.11]; K10788, eosinophil peroxidase[EC:1.11.1.7];K19511, peroxidase[EC:1.11.1.7];K20205, manganese peroxidase[EC:1.11.1.13];K03781,catalase[EC:1.11.1.6]

Fig.3 Differentially expressed gene(DEG)distributions in regenerated white oak sprouts of different ages.a Up-and downregulated DEG numbers in pairwise comparisons. “Up”in A:B comparisons indicates upregulated in B.“Down”in A:B comparisons indicates downregulated in B.b, c DEG distributions in leaves(b)and shoots(c). Horizontal bars indicate the DEG number in one comparison;vertical bars indicate DEG numbers shared in several comparisons
The number of DEGs in shoots was an order of magnitude lower than in leaves, with only nine pathways varying across ages, and no altered pathways were found in the S5:S40 comparison (Table 2 and S2). Half the DEGs in the starch and sucrose metabolism pathway were downregulated in older shoots, while the other half were upregulated. This trend was unlike that in leaves(Fig. 4b). The plant-pathogen interaction pathway changed with sprout growth. The majority of the DEGs in this pathway were downregulated at the 5a-10a stage but upregulated at the 10a-20a and 20a-40a stages. Accordingly, to respond to pathogens, the phagosome pathway was upregulated at 20a. The MAPK signaling pathway, a senescence-related pathway (Zhang et al.2020), was upregulated at the 10a-20a and 20a-40a stages. The phagosome pathway was upregulated in the S20 group, and the plant hormone signal transduction pathway was upregulated in older shoots (Fig. 4b).
Genes were separated on the basis of expression profiles
In leaves, six gene expression profiles were identified during regenerated sprout growth (Fig. 5). The downtrending expression profiles (0/2 and 7) contained genes related to energy, sugar, lipid, amino acid and nitrogen metabolism, which decreased with sprout age. The uptrending expression profiles (10 and 17/19) contained genes related to DNA replication, DNA repair and signal transduction, which increased with sprout age. Expression profile 18 contained genes related to MAPK signal and chlorophyll metabolism that were highly expressed in the 20a group.
In shoots, five gene expression profiles were identified during regenerated sprout growth (Fig. 6). The downtrending expression profiles (0 and 2) contained genes related to transporter, amino acid metabolism and nitrogen metabolism, which decreased with sprout age. The up-trending expression profiles (12 and 17/19)contained genes related to signal transduction, plant-pathogen interactions and translation, which increased with sprout age. Expression profiles 11/18 contained genes related tothe ubiquitin system, cytochrome P450 and proteasome that were highly expressed in the 20a group.
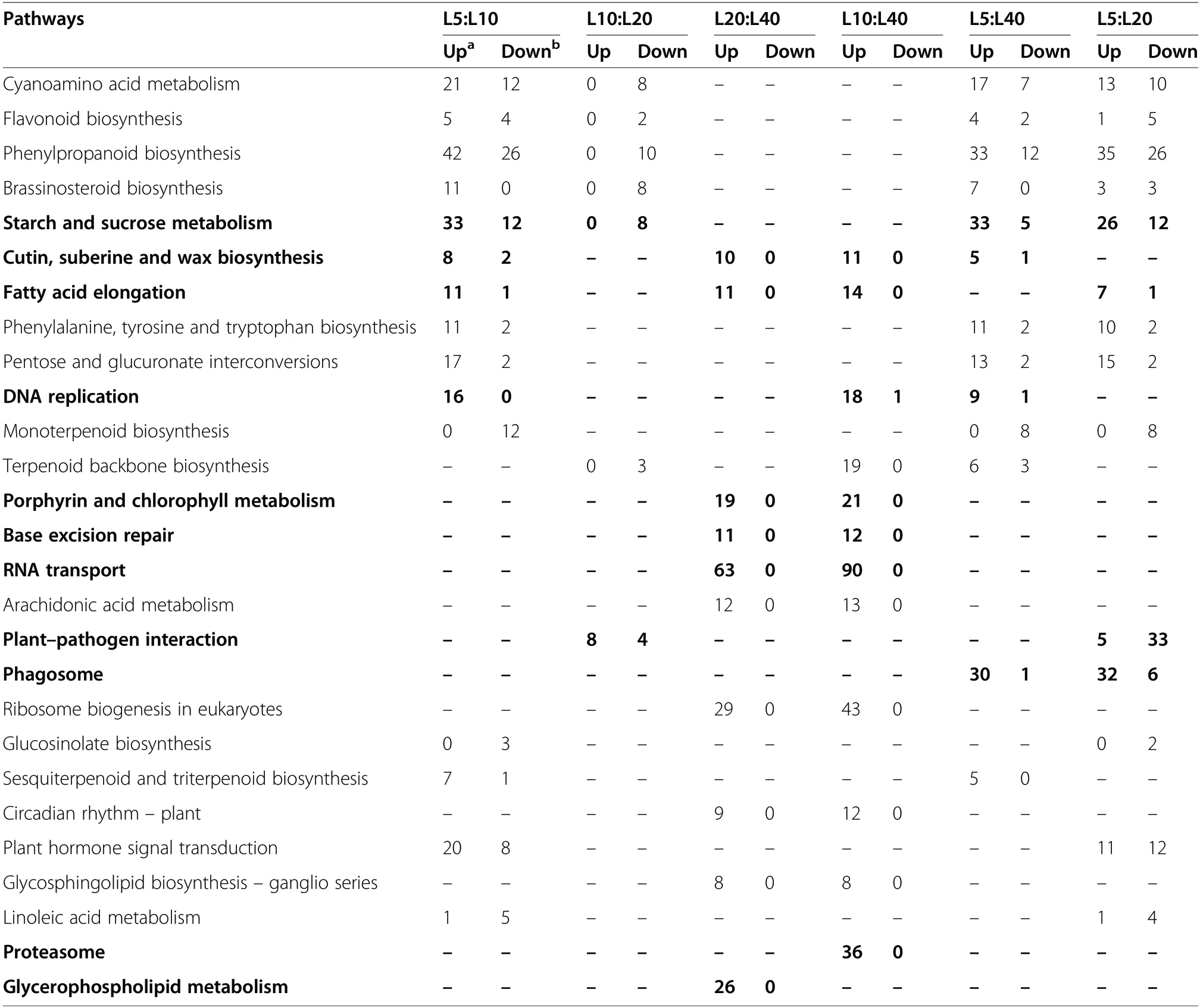
Table 1 KEGG enrichment of DEGs in leaves during regenerated white oak spssssrout growth(p<0.05)
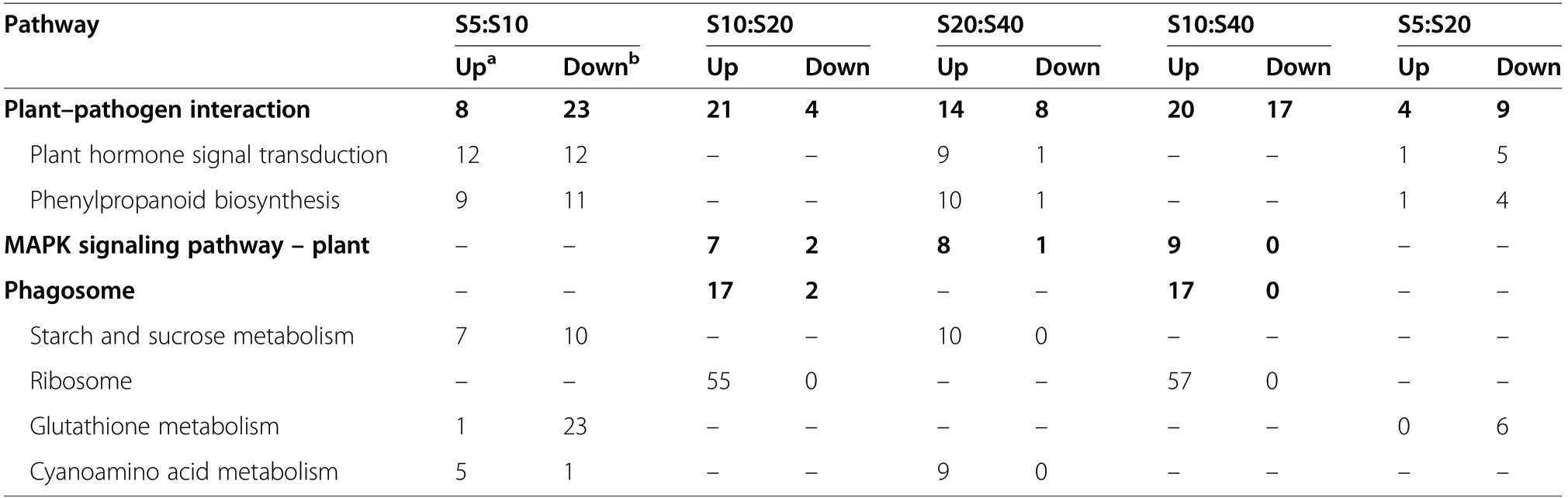
Table 2 KEGG enrichment of DEGs in shoots during regenerated white oak sprout growth(p<0.05)
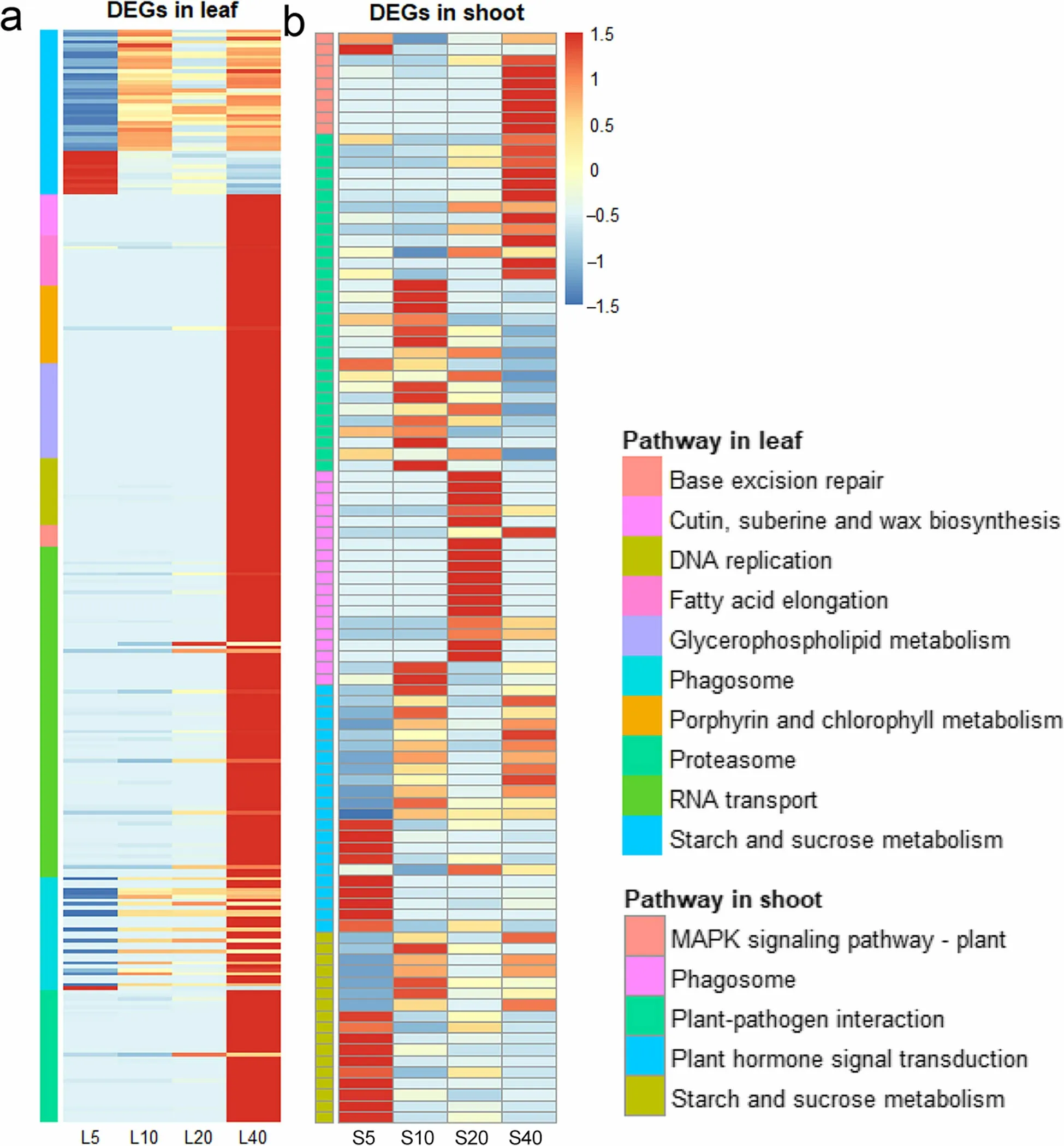
Fig.4 Expression levels of DEGs in pathways related to cell lifecycles in white oak leaves(a)and shoots(b). Data in this figure were normalized using z-scores
Discussion
Perennial plants grow continuously throughout their lives, with the meristem producing and rejuvenating new organs to compensate for those that are lost. The organism prevents senescence through continuous growth,which make it difficult to observe whole perennial plant senescence (Munné-Bosch 2015). The early aging of new meristematic products, that is, leaves and shoots, can indirectly prove plant senescence (Ryser et al. 2020). To ascertain whether regenerated sprouts succeed the parent’s age, which indicates that regenerated sprouts have shorter lifespans and age faster than sexually reproduced sprouts, the senescence levels of leaves and shoots over 40 years of growth were estimated. We did not find enough physiological or transcriptional evidence to support that regenerated sprouts senesce in 40 years.

Fig.5 Gene expression patterns in white oak leaves.Each box represents an expression profile.The upper and lower numbers in the boxes are the profile serial number and the p-value,respectively. Boxes of the same color represent similar expression profiles having high correlation rates.Bar plots represent the KEGG enrichment results for the genes in the profile,with the 20 most significant pathways being represented
Senescence is a programmed cell death phenomenon that involves oxidative stress, which is indicated by a decrease in antioxidative enzyme activity levels and peroxidation product accumulation. The peroxidation products lead to further DNA damage and cell membrane disruption (Juvany et al. 2013; Sade et al. 2018). In plants, respiration and photosynthetic activities are vigorous in summer. High levels of oxygen free radicals are produced in electron transport chains, and the antioxidative defense system is important in preventing oxidative damage (Juvany et al. 2013). Here, several antioxidative enzymes showed lower activities in older sprouts,but peroxidation products did not accumulate (Figs. 1 and 2).This indicated that the 35-year period only slightly influenced the antioxidative capabilities both at the transcriptional and physiological levels. The 40-year-old regenerated sprouts still maintained the capacity to eliminate peroxidation products.This trend was different from senescent plant organs. During Tulipa gesneriana petal senescence, antioxidative enzyme activity and ROS levels increase (Wang et al. 2020), while, ROS levels increased but antioxidative enzyme activity levels decreased during rice leaf senescence(Zhou et al.2020).
During cell senescence, macromolecules tend to be catabolized rather than synthesized; consequently, nutrients are transferred to healthy cells (Avila-Ospina et al.2014; Qi et al. 2020; Shinozaki et al. 2020). For example,during the bamboo shoot postharvest stage, pathways,including autophagy, fatty acid degradation and amino acid metabolism, are upregulated and accompanied by organism senescence (Li et al. 2019a, b). In this study,the main differences between ages were related to metabolism (Figs. 4, 5 and 6), with older regenerated sprouts having greater synthetic reactions in biosynthetic pathways, including flavonoid, brassinosteroid, wax, fatty acids and amino acids, compared with younger ones,and even the gene expression and cell division (RNA transport and DNA replication) levels increased with age(Table 1, Fig. 4a). These transcriptional changes indicate that 40-year-old sprouts still produce enough materials to maintain a large organism. However,the base excision repair pathway was upregulated in the L40 group, which indicates that there was more DNA damage in 40-yearold leaves (Li et al. 2020). Furthermore, the upregulation of phagosome, proteasome and glycerophospholipid metabolism suggested that tissue remodeling was accompanied by DNA damage in the L40 group (Fig. 4a).Although the autophagy pathway was not activated in the 40a group, the accumulation of DNA damage was an indicator of declining cell health, because it induces terminal programed cell death when the DNA damage exceeds the repair system’s threshold (Yamada and Coffman 2005; Martin and Wood 2019).
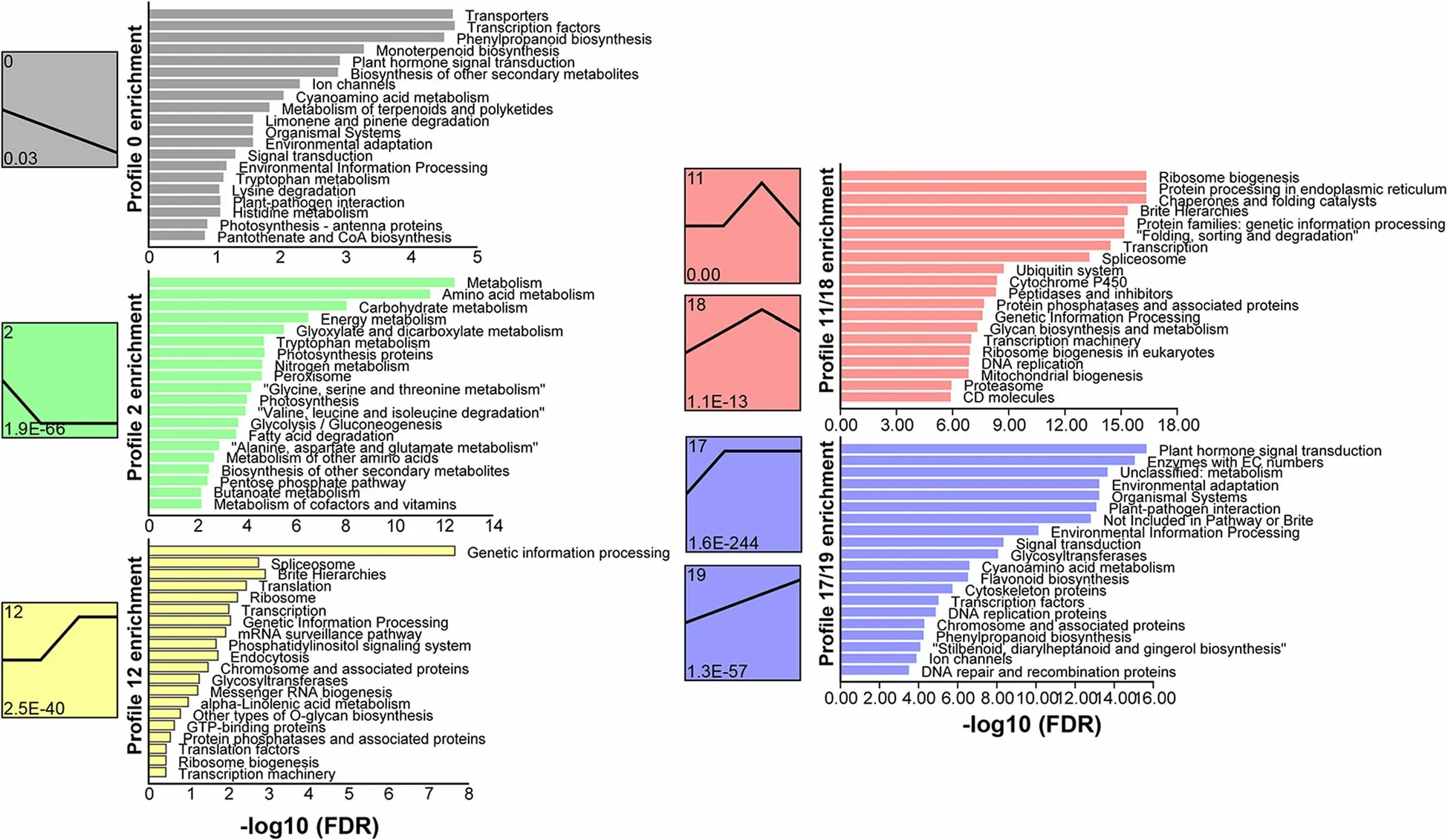
Fig.6 Gene expression patterns in white oak shoots.Each box represents an expression profile.The upper and lower numbers in the boxes are the profile serial number and the p-value,respectively. Boxes of the same color represent similar expression profiles having high correlation rates.Bar plots represent the KEGG enrichment results for the genes in the profile,with the 20 most significant pathways being represented
Unlike leaves, shoots act as transporters rather than producers, and they have longer lifespans in deciduous plants. Thus, strong metabolic activities are not required for shoot cell production and growth; consequently, the transcriptional differences in shoots were much smaller than in leaves. The main transcriptional variation was in the plant-pathogen interaction pathway, which changed with age (Table 2), which indicates that shoots faced age-related biotic stresses. Both effector-triggered immunity and pathogen associated molecular patterns(PMAP)-triggered immunity (PTI) were influenced by sprout age (Table S2). The PTI responses produce ROS to inhibit pathogen growth (Boller and Felix 2009), and the trends in shoot SOD and CAT activity levels were similar to the trend in the plant-pathogen interaction pathway. PTI responses activate the MAPK pathway(Meng and Zhang 2013), and in this study, the MAPK pathway was simultaneously upregulated in older shoots(Fig. 4b). MAPK signals induce events, including plant senescence, ROS generation and hypersensitive response,in Arabidopsis thaliana (Zhou et al. 2009; Matsuoka et al. 2015; Kawasaki et al. 2017).ducing lateral rosettes that share the same root, which results in the plant becoming polycarpic (Cotado and Munné-Bosch 2020). Lomatia tasmanica is an endangered species with only one population, and vegetative propagation is the only L. tasmanica reproduction method. The clonal age has been determined to be 43,46360000 years (Lynch et al. 1998). Thus, it appears that vegetative propagation and clonal growth prolong plant lifespans (Munne-Bosch 2014). Nevertheless, advanced clonal ages diminish or even delete the sexual propagation capacity, as seen in species such as L. tasmanica(Lynch et al. 1998), P. tremuloides (Ally et al. 2010) and Lemna minor (Barks and Laird 2015). Although these
In natural forests, sprouts are not artificially rejuvenated. Therefore, if regenerated sprouts inherit the parent’s age-related characteristics, then senescence symptoms should be more conspicuous. However, we did not find evidence that sprouts had inherited their parent’s age-related characteristics. This has been observed in other species. A 10-year phenological study in Brazil found that Diospyros lasiocalyx trees in three forests reproduced mainly through vegetative propagation,and widespread senescence in the D. lasiocalyx populations was not observed (Aguiar et al. 2020). Pyrenean saxifrage is a monocarpic perennial species that escapes monocarpic senescence at high altitudes by clonally prospecies still propagate vegetatively and are not senescent,the lack of sexual propagation decreases gene functional redundancy and population stress resistance.
Many forest tree species lose their regenerative capacities with age or maturation. Trees cut at more advanced ages have lower organ regenerative capabilities or regenerate organs having the parental characteristics resulting in low growth rates (Diaz-Sala 2014). However, a study reported that meristems from old-growth trees grafted onto juvenile rootstock are just as vigorous as those from mid-aged trees, although the former exhibited the old-tree parental characteristics (Greenwood et al. 2010).This indicated that meristem vigor does not decline with age and that the excepted lifespans of clonal trees are restrained by natural selection rather than genetic factors.Another study found that trees with a habitat-specific optimum body size have minimal mortality probability and maximal fertility in populations (Dani and Kodandaramaiah 2019). In this study, we hypothesized that two factors protected white oak sprouts from senescence, as follows: (1) sprouts were regenerated from stumps and their body size were smaller than parental,which indicated that the habitat was suitable for sprout growth and allowed the sprouts to absorb nutrients through the parental roots; and (2) callus cells reversed age-related effects during sprout formation, and consequently, the regenerated sprouts did not receive the parent’s age-related characteristics.
Undeniably, this study has some defects, as follows. (1)Technique-related limitations. The ages of sprout parents may not be consistent and errors in predicted ages may influence the sprout phenotypes. All the parental stumps had similar diameters, and the trees are from one population. The similar genetic background ensures that any age-related errors are not large. (2) The white oak lifespan is more than 400 years; therefore, a long time period is required for senescence symptoms to become obvious (Johnson and Abrams 2009). The results in this study indicate that 40-year-old sprouts are very healthy and are unlikely to senesce soon. (3) The experimental forest in the present study is located in a relatively comfortable environment and faces less abiotic and biotic stresses than other forests, such as windbreak forests and commercial plantations. The effects of stress resistance on regenerated sprouts have not been estimated; therefore, climate, interspecific competition and nutrient limitations may accelerate the aging of sprouts,especially clonal trees of the same genotype (Sloan and Sayer 2015; Zhang et al. 2018).
Conclusions
We found that in a natural forest, regenerated stump sprouts did not show negative signs of aging in the first 40 years of growth. Older sprouts still had robust antioxidative systems, while endocrine disorders, strong catabolism and massive autophagy were undetected,compared with younger sprouts. However, older sprouts had a few unhealthy characteristics, such as increased levels of DNA damage in leaf cells and more frequent tissue remodeling. Additionally, shoots suffered more biotic stress from pathogens, and MAPK signals were activated. The data indicated that although invisible injuries were accumulating, 40-year-old sprouts were still in the prime of their lives. The physiological age of parental trees did not significantly shorten the lifespans of the clonal offspring. Thus, although regenerated sprouts were not as strong as parental trees, they maintain the parental genotypes and help to sustain genetic diversity in harvested forests.
Supplementary Information
The online version contains supplementary material available at https://doi.org/10.1186/s40663-021-00292-1.
Additional file 1.
Additional file 2.
Abbreviations
ROS: Reactive oxygen species; MDA: Malondialdehyde; H2O2: Hydrogen peroxide; O2−:Superoxide anion; SOD: Superoxide dismutase;POD: Peroxidase; CAT: Catalase; KEGG: Kyoto Encyclopedia of Genes and Genomes; PTI: Pathogen associated molecular patterns (PMAP)-triggered immunity
Acknowledgements
We thank Lesley Benyon, PhD,from Liwen Bianji, Edanz Group China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Authors’contributions
Honggang Sun and Liwen Wu contributed to the conception and design of the study; Sisheng Wu organized the data and performed the statistical analysis; Honggang Sun and Sisheng Wu wrote the manuscript.All authors approved the submitted version.
Funding
This research was funded by the Fundamental Research Funds for the Central Non-profit Research Institution of CAF (CAFYBB2018ZB001).
Availability of data and materials
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Received: 23 August 2020 Accepted: 1 February 2021
- Forest Ecosystems的其它文章
- Bridging mapping and simulation modelling in the ecosystem service assessments of boreal forests:effects of bioenergy production on carbon dynamics
- Effects of three coniferous plantation species on plant-soil feedbacks and soil physical and chemical properties in semiarid mountain ecosystems
- Using GEDI lidar data and airborne laser scanning to assess height growth dynamics in fast-growing species:a showcase in Spain
- Decomposition and stabilization of organic matter in an old-growth tropical riparian forest:effects of soil properties and vegetation structure
- A multi-purpose National Forest Inventory in Bangladesh:design,operationalisation and key results
- Seamless integration of above-and undercanopy unmanned aerial vehicle laser scanning for forest investigation

